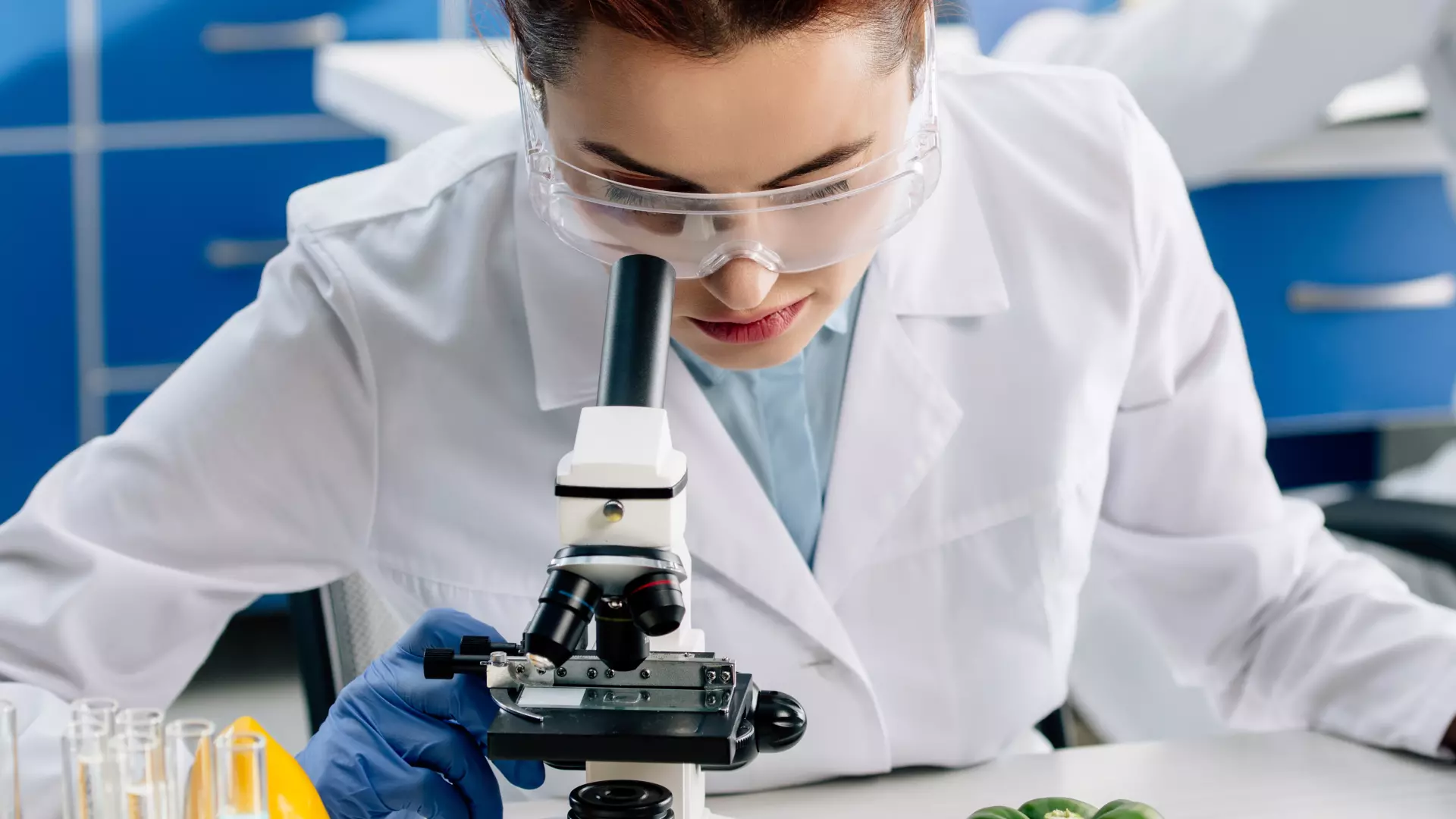

News
Do you want to stay up to date with the latest BHD news? This is the place to be!
Here we will let you know about the latest BHD research, events, and opportunities to get involved with our work.
Effective diagnosis for rare lung conditions
In this blog, we look at a recent review published by the UK Cystic Lung Disease Collaborative Network.

A Conversation with Dr Bryndis Yngvadottir
In this blog, we talk to Dr Bryndis Yngvadottir, a Research Associate in Medical Genetics at the University of Cambridge.

Pneumothorax at 17: My BHD Story
In a special piece to mark Rare Disease Day 2024, we speak to Kirsten. Kirsten shares her experiences of being diagnosed with a rare disease and things she learned along the way.
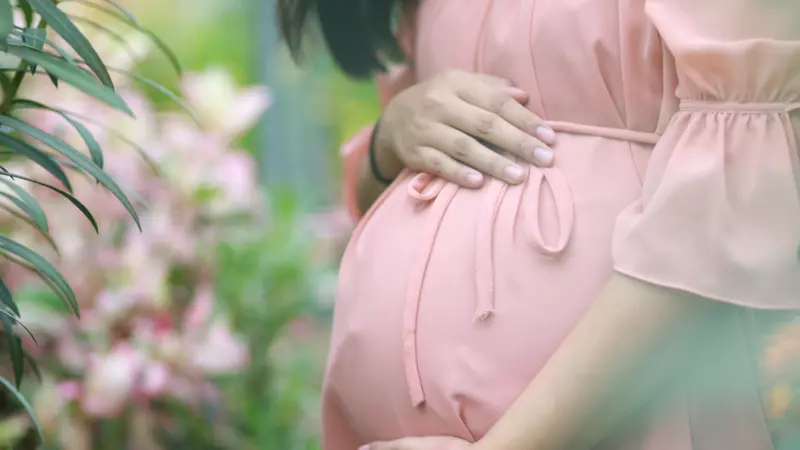

Pregnancy and Pneumothorax: What Does the Research Say?
In this blog, we take a look at a study that focuses on pregnancy and lung collapses.
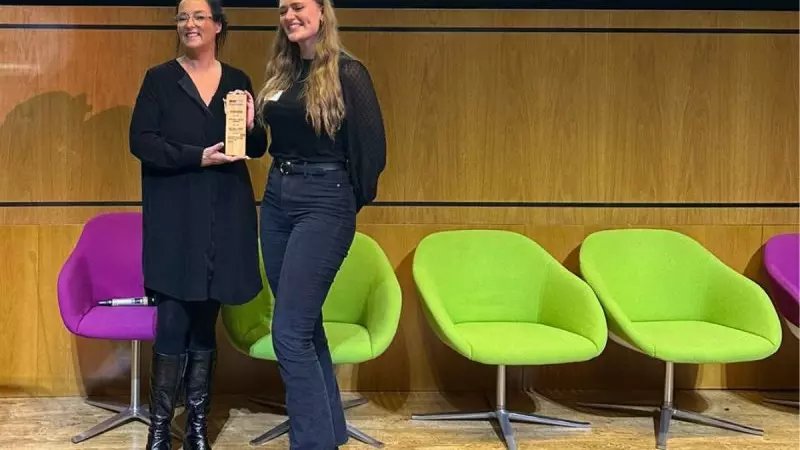

Early Career Profile: Rare Disease PhD Researcher Jennifer Heritz
In this blog, we talk to Jennifer about her goals for working in the rare disease research field, what she loves about her job and more.
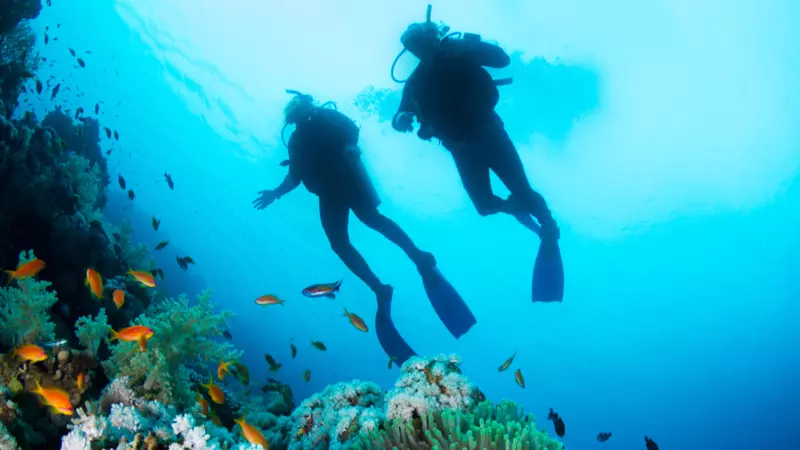

Is Scuba Diving Safe with Birt-Hogg-Dubé Syndrome?
Whilst scuba diving for many people can be a thrill-seeking experience, there are several risks to be considered if you have BHD, and it is important to be aware of and consider recommendations from clinical experts to keep yourself safe.
How lung cysts progress in BHD: A long term patient study monitoring l
December blog on How lung cysts progress in BHD: A long term patient study monitoring lung cyst growth

Pathogenic variants of FLCN and kidney cancer
Are there pathogenic variants of FLCN that do not cause kidney cancer, thus avoiding a requirement for lifelong surveillance? Read this blog to find out.
Who can be referred for genetic testing for inherited kidney cancer?
In this blog we talk about undergoing genetic risk assessment, how testing is performed and more.

Exploring a molecular link between BHD and Tuberous Sclerosis
Tuberous Sclerosis and Birt-Hogg-Dubé Syndrome share some similar features as well as manifestations that are unique to each condition. In this blog, we discuss more.
Is there a Beneficial Role for loss of FLCN in the Liver?
A study by has uncovered a surprising role for FLCN in the liver in the context of fatty liver disease progression. Read this blog to learn more.

Prevalence of BHD – an Epidemiological Study
To determine the prevalence of BHD, researchers from the University of Lausanne conducted a meta-analysis of previously published studies on the prevalence of spontaneous pneumothorax among BHD patients and the general population.